SNAP-CUTANA™HA Tag Panel提供了用于验证抗HA抗体和确认涉及HA表位标记的染色质蛋白的成功CUT&RUN反应的检测控制��。这一重要的阳性对照指导排除故障����,以区分HA表位标签问题(包括转基因表达��、标记蛋白的染色质结合����、标签的溶剂可及性等)与CUT&RUN工作流程中的技术故障����。该面板由两个包含未修饰组蛋白H3或3xHA-H3融合的核小体组成���,每个核小体都包裹有两个独特的条形码DNA模板(A和B���,用于内部技术复制)��。核小体分别与顺磁珠偶联��,并汇集成一个Panel����,方便一步插入到切割和运行反应�����。在添加抗ha或IgG阴性对照抗体之前���,将该检测组合与ConA固定的细胞一起添加(见应用说明和表1)���。pAG-MNase释放基因组染色质和条形码核小体取决于所用抗体的特异性����。测序后�����,恢复的HA与未修饰的核小体的相对读长计数提供了在靶恢复与脱靶恢复的定量指标(图2)���,从而衡量实验的成功率��,并指导故障排除工作���。有关工作流集成����、预期结果��、数据分析和故障排除的详细信息��,请参阅最新的CUTANA™ CUT&RUN方法(https://www.epicypher.com/resources/protocols/cutana-cut-and-run-kit-manual)和SNAP-CUTANA™ Spike-in(https://www.epicypher.com/resources/protocols/)用户指南����。
保存温度
自收到之日起��,-20℃下可稳定储存6个月���。较低的温度会导致冻结���,并会永久损坏磁珠��。
验证数据
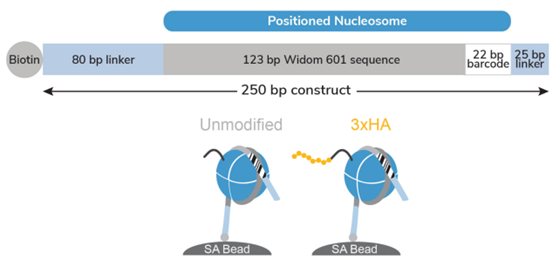
Figure 1: Schematic of SNAP-CUTANA™ HA Tag Panel |
The HA Tag Panel contains two nucleosomes - one has an H3 tail fusion to a 3xHA Tag epitope and one is an unmodified control. Both octamers are wrapped with two uniquely barcoded DNA templates (A and B). Each 250 bp DNA template contains a 123 bp 601 nucleosome positioning sequence (gray) [1], a unique 22 bp DNA-barcode (white; 4 barcodes total), and a 5’ biotin-TEG. The 5’ and 3’ linkers (blue) are compatible with cleavage by pAG-MNase (EpiCypher 14-1048, 15-1016) during CUT&RUN. The nucleosomes are individually pre-conjugated to paramagnetic beads and pooled for convenient use.
|
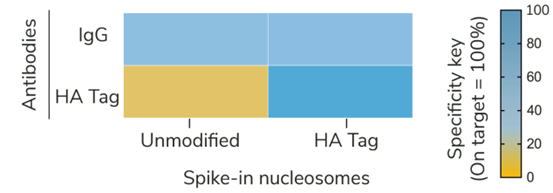
Figure 2: SNAP-CUTANA™ HA Tag Panel provides an in-assay control for CUT&RUN reactions targeting HA-tagged proteins |
CUT&RUN was performed as described in Figure 5. CUT&RUN sequencing reads were aligned to the unique DNA barcodes corresponding to each nucleosome in the SNAP-CUTANA™ HA Tag Panel. Data are expressed as a percent relative to on-target recovery (HA Tag set to 100%) or total counts (IgG). IgG antibody results demonstrate equal loading of unmodified and epitope nucleosomes in the panel. HA Tag antibody results show selective enrichment of the HA Tag spike-in nucleosomes, validating all CUT&RUN steps, including HA antibody binding, pAG-MNase cleavage, and wash conditions
|
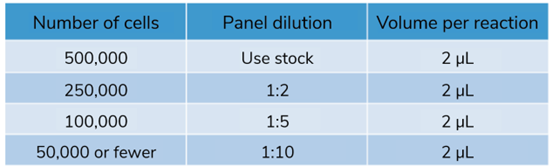
Table 1: Recommended SNAP-CUTANA™ HA Tag Panel Spike-in dilution for CUT&RUN reactions of varying starting cell number. |
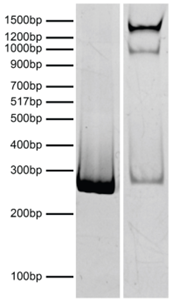
Figure 3: DNA gel data |
Nucleosomes in the SNAP-CUTANA™ HA Tag Panel were resolved via native PAGE and stained with ethidium bromide to confirm intact nucleosome assembly. Lane 1: Free 250 bp DNA used in nucleosome assembly (100 ng). Lane 2: Intact nucleosomes (400 ng).
|
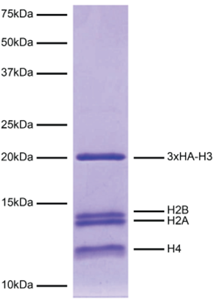
Figure 4: Protein gel data |
Coomassie stained SDS-PAGE gel of the nucleosome containing a 3xHA-H3 fusion (1 μg) in the SNAP-CUTANA™ HA Tag Panel demonstrates the purity of histones in the preparation. Sizes of molecular weight markers and positions of the core histones (H2A, H2B, 3xHA-H3, and H4) are indicated.
|
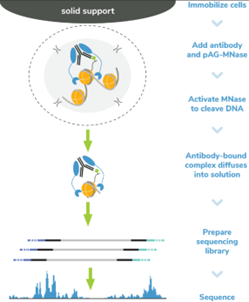
Figure 5: CUT&RUN methods |
CUT&RUN was performed on 500k MDA-MB-231 native cells stably expressing 3xHA-tagged GATA3 [1]* using the CUTANA™ ChIC/CUT&RUN Kit v3 (EpiCypher 14-1048). SNAP-CUTANA™ HA Tag Panel was added just prior to the addition of either HA Tag (0.5 µg; EpiCypher 13-2010) or IgG negative control (0.5 µg; EpiCypher 13-0042) antibodies. Library preparation was performed with 5 ng of DNA (or the total amount recovered if less than 5 ng) using the CUTANA™ CUT&RUN Library Prep Kit (EpiCypher 14-1001/14-1002). Libraries were run on an Illumina NextSeq2000 with paired-end sequencing (2x50 bp). Data were aligned to the hg19 genome using Bowtie2. Data were filtered to remove duplicates, multi-aligned reads, and ENCODE DAC Exclusion List regions.
*Thanks to Dr. Takaku (UND) for 3xFLAG-GATA3-3xHA MDA-MB-231 cells. |
订购详情
货号 | 产品名称 | 规格 |
19-5002 | SNAP-CUTANA™ HA Tag Panel | 50 Reactions |
参考文献
[1] Lowary & Widom J. Mol. Biol. (1998). PMID: 9514715
[2] Takaku et al. Genome Biol. (2016). PMID: 26922637

如需了解更多详细信息或相关产品����,
请联系EpiCypher中国授权代理商-欣博盛生物
全国服务热线: 4006-800-892 邮箱: market@hnyazd.com
深圳: 0755-26755892 北京: 010-88594029
广州��:020-87615159 上海: 021-34613729
代理品牌网站: www.hnyazd.com
自主品牌网站: www.neobiosescience.ne